Chromosome entrapment assay
Chromosome entrapment assay
Question: Does a 'sliding clamp' protein entrap chromosomal DNA?
Approach: immobilize putative chromosome-bound clamp to chromosome, assess if infact bound
I. Engineer protein of interest for optimal cross-linking and detection
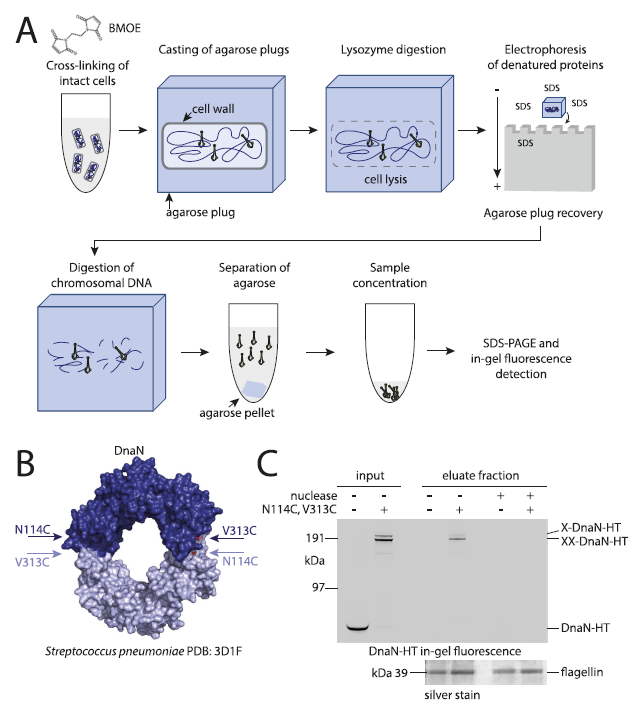
Bi) Engineer cysteine residues into DnaN (replicative sliding clamp) inorder to cross-link into 'covalent rings' by cysteine-specific crosslinking agent (BMOE)
ii) Engineer C-term HaloTag to DnaN that can bind a fluorescent ligand for detection following experiment (DnaN-HT)
II. Subject engineered protein to assay
Ai) Cross-link cells with cysteine-specific cross-linker
Aii) Embed bacterial cells in agarose 'plug', then treat with lysozyme to digest cell walls (how does enzyme work on cells embedded in agarose?)
Aiii) Subject agarose plugs to electric field in presence of SDS-buffer to denature/remove unbound proteins (So DNA remains in plug following electrophoresis?)
- So even cross-linked proteins released while protein 'rings' physically retain association here?
Aiv) Treat agarose plugs with nuclease to digest away genomic DNA
Av) Pellet agarose, retain eluate with released proteins
Avi) concentrate protein, resolve via SDS-PAGE
Avii) visualize via HaloTag ligand or silver stain (why not WB)?
III. Visualize and interpret
Input fractions:
Ci) Input lane 1: Non-cross-linked Dna-HT deteced
Input lane 2) top band - single crosslinked DnaN. bottom band - double cross-linked DnaN
Cii) Eluate fractions
Eluate lane 1: lack of engineered cross-linker sites does not retain DnaN-HT
Eluate lane 2: presence of engineered cross-linker sites retains DnaN-HT in assay
Eluate lanes 3-4: nuclease treatment reduces retention of entrapped proteins (but why would this happen? Are proteins pelleted with fragments?)
Approach: immobilize putative chromosome-bound clamp to chromosome, assess if infact bound
I. Engineer protein of interest for optimal cross-linking and detection
Bi) Engineer cysteine residues into DnaN (replicative sliding clamp) inorder to cross-link into 'covalent rings' by cysteine-specific crosslinking agent (BMOE)
ii) Engineer C-term HaloTag to DnaN that can bind a fluorescent ligand for detection following experiment (DnaN-HT)
II. Subject engineered protein to assay
Ai) Cross-link cells with cysteine-specific cross-linker
Aii) Embed bacterial cells in agarose 'plug', then treat with lysozyme to digest cell walls (how does enzyme work on cells embedded in agarose?)
Aiii) Subject agarose plugs to electric field in presence of SDS-buffer to denature/remove unbound proteins (So DNA remains in plug following electrophoresis?)
- So even cross-linked proteins released while protein 'rings' physically retain association here?
Aiv) Treat agarose plugs with nuclease to digest away genomic DNA
Av) Pellet agarose, retain eluate with released proteins
Avi) concentrate protein, resolve via SDS-PAGE
Avii) visualize via HaloTag ligand or silver stain (why not WB)?
III. Visualize and interpret
Input fractions:
Ci) Input lane 1: Non-cross-linked Dna-HT deteced
Input lane 2) top band - single crosslinked DnaN. bottom band - double cross-linked DnaN
Cii) Eluate fractions
Eluate lane 1: lack of engineered cross-linker sites does not retain DnaN-HT
Eluate lane 2: presence of engineered cross-linker sites retains DnaN-HT in assay
Eluate lanes 3-4: nuclease treatment reduces retention of entrapped proteins (but why would this happen? Are proteins pelleted with fragments?)