Transposable Elements
All mobile DNA elements are classified as transposable elements (TEs). Active TE's attempt to increase copy number at the cost of host fitness due to mutagenic effects of genome re-integration, including DDR activation from excessive dsDNA breaks, genome rearrangement, chromosome breakage, alteration of slicing or polyadenylation patterns, and serving as de novo promoters or enhancers. To combat the harmful effects of active TE's, the genome has evolved overlapping epigenetic 'defense' mechanisms that silence the expression and mobility of transposable elements.
Full-length autonomous TEs encode all proteins required for transposition, but remain silent as cryptic elements epigenetically silenced by the genome. While TE distribution throughout the genome appears indiscriminate, cryptic and non-autonomous TEs accumulate in heterochromatic regions of the genome, particularly at pericentromeric and telomeric regions where they are utilized by the genome in structural organization to regulate gene expression
Types and Structures of Transposable Elements
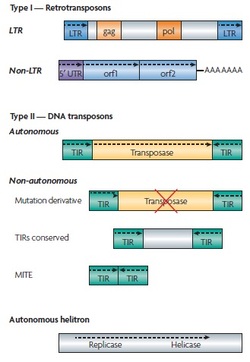
Type I elements - Retrotransposons (require reverse reverse-transcription step during duplicative transposition which increases total copy number)
L1 ORF2 encodes an endonuclease that nicks host DNA and a reverse transcriptase to synthesize L1 cDNA.
Type II Elements - DNA Transposons (cut and paste transposition)
Transposable Element Age
Full-length autonomous TEs encode all proteins required for transposition, but remain silent as cryptic elements epigenetically silenced by the genome. While TE distribution throughout the genome appears indiscriminate, cryptic and non-autonomous TEs accumulate in heterochromatic regions of the genome, particularly at pericentromeric and telomeric regions where they are utilized by the genome in structural organization to regulate gene expression
Types and Structures of Transposable Elements
Type I elements - Retrotransposons (require reverse reverse-transcription step during duplicative transposition which increases total copy number)
- LTR retrotransposons - flanked by LTRs, encode gag and pol proteins related to retroviral proteins, lack envelope protein required to leave cell
- Non-LTR retrotransposons - ex. LINE1 (L1) family, SINE elements (ex. Alu, SVA elements), etc.
L1 ORF2 encodes an endonuclease that nicks host DNA and a reverse transcriptase to synthesize L1 cDNA.
Type II Elements - DNA Transposons (cut and paste transposition)
- Encode transposase that recognizes terminal inverted repeats (TIRs) that flank the TE, excise element, intergrates it into new 'acceptor' site
- Gap left at original 'donor' site repaired without element, or filled with copy of transposon
Transposable Element Age
Silence of the Elements: Epigenetic Strategies
The genome employs diverse epigenetic defense mechanisms to suppress TE transcription, including modification on histone tails, DNA methylation, and alterations in chromatin packing and condensation.
Histone Modifications
Nucleosomes associated with TEs are enriched for H3K9 methylation which marks transcriptionally repressive and inactive chromatin. Inactivation of the H3K9 methyltransferase Suv39 results in modest upregulation of TE transcripts in mouse embryonic stem cells.
DNA Methylation
Inheritable DNA methylation marks on cytosine residues (CpG) provide another mechanism of TE silencing. In mice, the DNMT1 methyltransferase maintains DNA methylation. In DNMT1-deficient mice (embryonic lethal E9-10) elevated intracisternal A-particle (IAP) retrotransposon transcript levels are detected in embryos, but not undifferentiated ES cells. DNMT3 establishes de novo methylation in a stage-specific manner during embryogenesis. Ablation of the non-catalytic homologue DNMT3L results in IAP retrotransposon expression due to disruption of methylation in premeiotic male germ cells.
Modification of Chromatin Structure
SWI/SNF chromatin remodelling proteins utilize ATP hydrolysis to remodel nucleosomes and silence TEs. In mice, ablation of the the SWI2/SNF2-like protein Lsh1 results in loss of DNA methylation in embryos, and TE expression.
Histone Modifications
Nucleosomes associated with TEs are enriched for H3K9 methylation which marks transcriptionally repressive and inactive chromatin. Inactivation of the H3K9 methyltransferase Suv39 results in modest upregulation of TE transcripts in mouse embryonic stem cells.
DNA Methylation
Inheritable DNA methylation marks on cytosine residues (CpG) provide another mechanism of TE silencing. In mice, the DNMT1 methyltransferase maintains DNA methylation. In DNMT1-deficient mice (embryonic lethal E9-10) elevated intracisternal A-particle (IAP) retrotransposon transcript levels are detected in embryos, but not undifferentiated ES cells. DNMT3 establishes de novo methylation in a stage-specific manner during embryogenesis. Ablation of the non-catalytic homologue DNMT3L results in IAP retrotransposon expression due to disruption of methylation in premeiotic male germ cells.
Modification of Chromatin Structure
SWI/SNF chromatin remodelling proteins utilize ATP hydrolysis to remodel nucleosomes and silence TEs. In mice, ablation of the the SWI2/SNF2-like protein Lsh1 results in loss of DNA methylation in embryos, and TE expression.
Transposable Elements affect chromosome structure and function
Centromeres and telomeres are repeat-rich regions of chromosomes composed primarily of constitutive heterochromatin. With respect to chromosome function and genome integrity, centromeres are essential in chromosome trafficking while telomeres prevent chromosome shortening following replication.
Centromeric Regions
Centromeres consist of long tandem arrays of simple sequence repeats called satellite repeats, surrounded by a peripheral region rich in TEs. Reactivation of epigenetically silenced satellite repeats and TEs is accompanied by chromosome segregation and meiotic defects in mice, loss of chromatid cohesion in fission yeast, and loss of centromere condensation in A. thaliana.
Telomeres
sssss
Centromeric Regions
Centromeres consist of long tandem arrays of simple sequence repeats called satellite repeats, surrounded by a peripheral region rich in TEs. Reactivation of epigenetically silenced satellite repeats and TEs is accompanied by chromosome segregation and meiotic defects in mice, loss of chromatid cohesion in fission yeast, and loss of centromere condensation in A. thaliana.
Telomeres
sssss
Transposable Elements in Cancer
Almost all human cancers exhibit genome-wide hypomethlation of TE promoters associated with elevated TE expression, and loss of critical DNA sensing and repair pathways. TE's can cause or exacerbate mutagenic genomic rearrangements that facilitate genomic instability in cancer.
Elevated full-length L1 mRNA expression is observed in the majority of human cancers and cancer-derived cell lines. Elevated expression is primarily attributed to aberrant DNA methylation which is disrupted early in the transformation process.
Insertional Mutagenesis
Transposable elements have evolved means to continue to mobilize, in particular within germ cells, as a means to ensure continued existence within the host. within
In general, it has been found that cells that repress apoptosis, or cells that have defects in certain DNA repair pathways support higher retrotransposition.
Human retroelements (L1, Alu, and SVA) aplify in the host genome by using RNA intermediates which leave the parental copy intact. Often, insertion occurs in genomic regions with crucial biological functions, resulting in insertional mutagenesis. Several examples of human cancers caused by Alu- and L1- mediated insertional mutagenesis exist. For example:
BRCA - breast cancer
APC - colon cancer
CRB1 - retinoblastoma (protein localizes to mammalian retina)
MYC - breast cancer
Recombination
Retroelements serve as potential sources of homology required for non-allelic homologous recombination (NAHR) (preferentially involve Alu elements). Consequences of NAHR are DNA damage that leads to duplications or deletions between the two participating sequences. Presence of Alu sequences in intronic regions promote 40-300x more recombination-induced deletions in mutant p53 backgound than WT. In tumorigenesis-prone backgrounds, repetitive elements increase recombination frequency and contribute to genomic instability. Several examples of human cancers caused by Alu/Alu NAHR enclude:
von-Hippel-Lindau tumor suppressor (VHL) gene - VHL syndrome (germ-line)
human caspase-activated DNase gene - heptoma (germ-line)
Re-arrangements both germ-line and somatic. Somatic Alu/Alu re-arrangements demonstrated via:
Duplication of MYB locus - T-cell acute lymphoblastic leukemia (T-ALL)
Duplication of exons in MLL (a histone methyl transferase) - acute myeloid leukemia (AML)
DNA Double strand breaks
Expression of L1 ORF2 generates DSBs 10-100x higher than the number of L1 retrotransposition events in HeLa cells. In the presence of such DNA lesions, checkpoints which normally prevent genome replication in the presence of unrepaired DNA damage are lost, allowing cells to progress through the cell cycle.
TE expression is not limited to genomic context and programmed responses. Expression and activity can also be stimulated by external stimuli. For example, etoposide treatment increases Alu retrotransposition rates. Cells exhibit elevated Alu expression following heat shock, or treatments with puromycin, cyclohexamide, etoposide and cisplati, exposure to UV radiation, or infection with Adenovirus type 2.
p53 utilizes p53 consensus sequences within the L1 promoter to activate L1 transcription, presumably to introduce more DSBs and amplify the p53-mediated DNA damage response.
Elevated full-length L1 mRNA expression is observed in the majority of human cancers and cancer-derived cell lines. Elevated expression is primarily attributed to aberrant DNA methylation which is disrupted early in the transformation process.
Insertional Mutagenesis
Transposable elements have evolved means to continue to mobilize, in particular within germ cells, as a means to ensure continued existence within the host. within
In general, it has been found that cells that repress apoptosis, or cells that have defects in certain DNA repair pathways support higher retrotransposition.
Human retroelements (L1, Alu, and SVA) aplify in the host genome by using RNA intermediates which leave the parental copy intact. Often, insertion occurs in genomic regions with crucial biological functions, resulting in insertional mutagenesis. Several examples of human cancers caused by Alu- and L1- mediated insertional mutagenesis exist. For example:
BRCA - breast cancer
APC - colon cancer
CRB1 - retinoblastoma (protein localizes to mammalian retina)
MYC - breast cancer
Recombination
Retroelements serve as potential sources of homology required for non-allelic homologous recombination (NAHR) (preferentially involve Alu elements). Consequences of NAHR are DNA damage that leads to duplications or deletions between the two participating sequences. Presence of Alu sequences in intronic regions promote 40-300x more recombination-induced deletions in mutant p53 backgound than WT. In tumorigenesis-prone backgrounds, repetitive elements increase recombination frequency and contribute to genomic instability. Several examples of human cancers caused by Alu/Alu NAHR enclude:
von-Hippel-Lindau tumor suppressor (VHL) gene - VHL syndrome (germ-line)
human caspase-activated DNase gene - heptoma (germ-line)
Re-arrangements both germ-line and somatic. Somatic Alu/Alu re-arrangements demonstrated via:
Duplication of MYB locus - T-cell acute lymphoblastic leukemia (T-ALL)
Duplication of exons in MLL (a histone methyl transferase) - acute myeloid leukemia (AML)
DNA Double strand breaks
Expression of L1 ORF2 generates DSBs 10-100x higher than the number of L1 retrotransposition events in HeLa cells. In the presence of such DNA lesions, checkpoints which normally prevent genome replication in the presence of unrepaired DNA damage are lost, allowing cells to progress through the cell cycle.
TE expression is not limited to genomic context and programmed responses. Expression and activity can also be stimulated by external stimuli. For example, etoposide treatment increases Alu retrotransposition rates. Cells exhibit elevated Alu expression following heat shock, or treatments with puromycin, cyclohexamide, etoposide and cisplati, exposure to UV radiation, or infection with Adenovirus type 2.
p53 utilizes p53 consensus sequences within the L1 promoter to activate L1 transcription, presumably to introduce more DSBs and amplify the p53-mediated DNA damage response.
